|
|
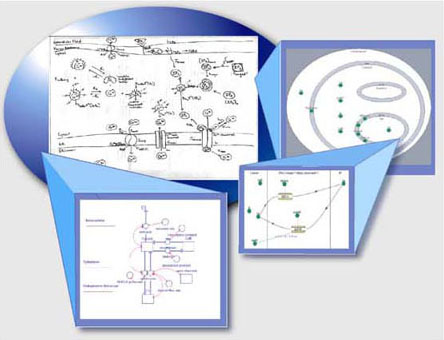
Tools
used during the workshop |
| Stella
| JDesigner | Excel
| Virtual Cell |
| Stella |
|
| This
software uses the metaphor of flows and stocks for
drafting models of dynamic systems. Model examples
and tutorials can be downloaded from the company site.
This
software is useful for Mapping and Modeling, Simulation
and Analysis, and Communication
|
|
|
| JDesigner |
|
This
application is used to draw biochemical networks and
export the drawn network as SBML (Systems Biology
Markup Language). It can be used as an interface for
developing networks that run on Jarnac.
|
|
|
|
|
|
Users
create models through a graphical user interface or
by generating mathematical descriptions using Virtual
Cell Math Language. The tool is accessible via the
web with user registration.
|
|
Other
tools not covered in the workshop but referenced by
students |
|
|
|
| This
is a tool for inferring and interpreting phylogenetic
trees, available for purchase at the company
website.
|
|
|
|
|
| This
is a free inference tool for phylogenetic analysis.
|
|
|
|
|
| The
main features underlying GenMAPP are:
- Draw pathways with easy to use graphics tools
- Color genes on MAPP files based on user-imported
genomic data
- Query genomic data against MAPPs and the GeneOntology
with MAPPFinder
- excerpted from the GenMapp
site.
|
|
|
|
|
| BRB
ArrayTools is an integrated package for the visualization
and statistical analysis of DNA microarray gene expression
data.
- excerpted from the BRB
array site.
|
|
|
|
|
| Collaborative
information management.
|
|
|
|
|
This
modeling framework makes uses a spreadsheet interface,
solves differential equations, allows users to enter
their own rate constants and input conditional statements
for the simulation.
Because
the cell cycle underlies the growth, development,
and reproduction of all living organisms, knowledge
about its control is central to cell biology and has
potential applications in the health care and pharmaceutical
industries. We want to develop novel, computational
tools, with user-friendly interfaces, for studying
complex biochemical regulatory systems in general,
and the cell cycle control system in particular.
- exerpted from the JigCell
site.
Paper on the JigCell Model Builder (PDF) |
|
| |